Network modules/communities
Description:
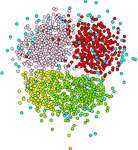
ModuLand:
Analysis of complex, real-world networksup to several million nodes, identification of key nodes and prediction of behavior.
Our work focuses on special methods for analysing the fine structure of complex networks comprising many interacting elements with directed or undirected links. In these networks the weight of a link is characterised by the strength of the interaction between its elements. Our investigations include methods for high resolution modularization of networks as well as methods for identifying distinguished links or elements supposedly playing special roles in the network (such as links or elements situated in module centers or overlaps).
Our family of methods for network analysis is based on rendering a novel index to all elements and links of the network called ‘community landscape height value’ quantitatively characterising the centrality of each element and link from the viewpoint of the network as a whole. We analise technical and social networks as well as protein networks.
Publications:
- Szalay-Bekő, M., Palotai, R., Szappanos, B., Kovács, I.A., Papp, B. and Csermely, P. (2012) ModuLand plug-in for Cytoscape: extensively overlapping modules, community centrality and their use in biological networks. Bioinformatics, in press http://arxiv.org/abs/1111.3033
Download it!
The Cytoscape plug-in program and its User Guide can be downloaded from here: http://www.linkgroup.hu/modules.php - Farkas, I.J., Korcsmáros, T., Kovács, I.A., Mihalik, Á., Palotai, R., Simkó, G.I., Szalay, K.Z., Szalay-Bekő, M., Vellai, T., Wang, S. and Csermely, P. (2011) Network-based tools in the identification of novel drug-targets. Science Signaling 4, pt3. Download the paper!

Download the slideshow! - Palotai, R. and Csermely, P. (2009) Network modules help the identification of key transport routes, signaling pathways in cellular and other networks. Annalen der Physik 18, 822-829, www.arxiv.org/0908.4524
Download it!
- Kovács, I.A., Palotai, R., Szalay, M.S. and Csermely, P. (2010) Community landscapes: a novel, integrative approach for the determination of overlapping network modules. PLoS ONE 7, e12528, IF: 4.4 Download it!

www.arxiv.org/abs/0912.0161 Supporting web-site with all downloadable algorithms: www.linkgroup.hu/modules.php
Featured in the February 2011 issue of Science Signaling - Csermely, P. (2008) Creative elements: network-based predictions of active centres in proteins, cellular and social networks. Trends Biochem. Sci. 33, 569-576, Download it!

Visit it at arXiv.org!
- Csermely, P., Korcsmáros, T., Kovács, I. and Szalay, M. (2006) Method for analyzing the fine structure of networks. Patent application WO2007093960.
- Palotai, R. Szalay, M.S. and Csermely, P. (2008) Chaperones as integrators of cellular networks: changes of cellular integrity in stress and diseases. IUBMB Life 60, 10-18, arxiv.org/0710.1622, IF: 2.3 Download it!

Visit it at arXiv.org! - Böde, C., Kovács, I.A., Szalay M., Palotai, R., Korcsmáros, T. and
Csermely, P. (2007) Network analysis of protein dynamics. FEBS Lett. 581
(15): 2776-82. IF: 3,5 Download it!

- Korcsmáros, T., Kovács, I. A., Szalay, M. S. and Csermely, P (2006) Molecular chaperones: The modular evolution of cellular networks. Journal of Bioscience 32 (3): 441-446. IF: 1,5 Download it!


Questions | Address | Design and maintenance | Visitors since 7th July 2004:
